|
Network and Systems Biology
Our unit works hand in hand with wet lab experts and it is often challanged to analize different kinds of data. Our Molecular Genetics Laboratory does classical experiments as well as high-throughput experiments using instruments such as automatic confocal microscopy, Luminex and CyTOF2. Thus, systems-level analysis are a key part of our work. We are focused on complex network analysis, graph theory, boolean networks and dynamical models design and manipulation using CellDesigner and COPASI. Our work is aimed to support further wet studies which in turn demand for more analysis.
Network analysis seeks to understand the relationships within biological networks such as metabolic or protein-protein interaction networks.
Although biological networks can be constructed from a single type of molecule or entity (such as genes), network biology often attempts to integrate many different data types, such as proteins, small molecules, gene expression data, and others.
Systems biology involves the use of computer simulations of cellular subsystems (such as the networks of metabolites and enzymes, signal transduction pathways and gene regulatory networks) to both analyze and visualize the complex connections of these cellular processes.
| |||
|
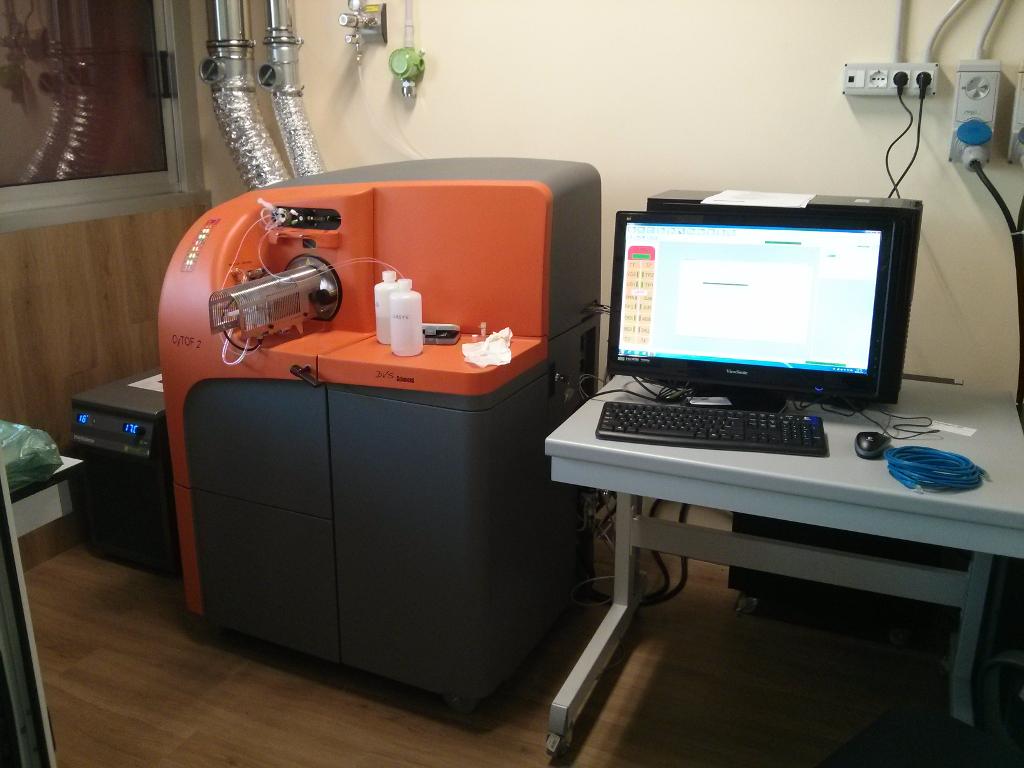
Mass cytometry (CyTOF2) is a mass spectrometry technique based on inductively coupled plasma mass spectrometry used for the determination of the properties of cells (cytometry). In this approach, antibodies are tagged with isotopically pure rare earth elements and these are used to tag the components of cells. Cells are nebulized and sent to an argon plasma, ionizing the multi-atom metal tags, which are then analyzed by a time-of-flight mass spectrometer. The approach overcomes limitations of spectral overlap that limit flow cytometry. [Wikipedia] | ||

|
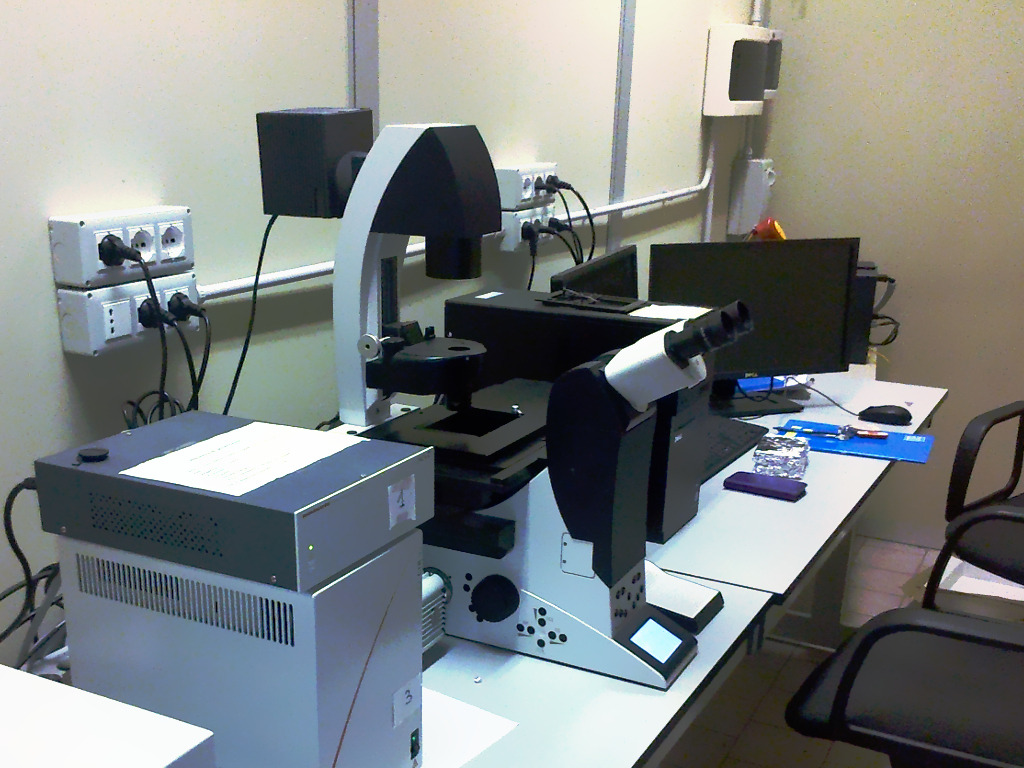
Confocal microscopy is an optical imaging technique for increasing optical resolution and contrast of a micrograph by means of adding a spatial pinhole placed at the confocal plane of the lens to eliminate out-of-focus light. It enables the reconstruction of three-dimensional structures from the obtained images. This technique has gained popularity in the scientific and industrial communities and typical applications are in life sciences, semiconductor inspection and materials science. [Wikipedia] | ||

|
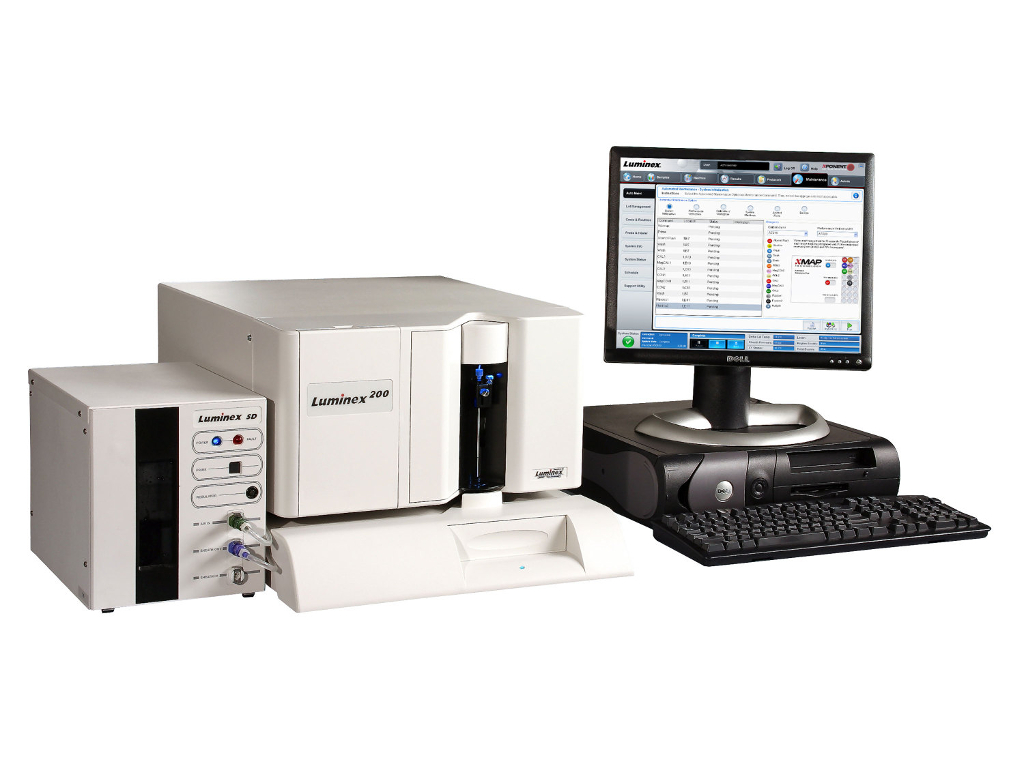
Luminex is bead based multiplexing, where beads are internally dyed with fluorescent dyes to produce a specific spectral address. Biomolecules (such as an oligo or antibody) can be conjugated to the surface of beads to capture analytes of interest. This technology uses flow cytometric or imaging technologies for characterization of the beads as well as detection of phycoerythrin emission due to analyte presence. The Luminex technology enables up to 500 proteins or genes to be detected in each well of a 96 or 384-well plate, using very small sample volume. Common applications include cytokines, metabolic markers, and phosphorylated proteins. [Wikipedia] | ||